What Sugar Must Be Added to the Growth Media to Ensure Expression of the Pmocab Operon?

The Influence of the Toxin/Antidote mazEF on Growth and Survival of Listeria monocytogenes under Stress
Department of Biotechnology and Biomedicine, Technical University of Kingdom of denmark, Matematiktorvet Bldg. 301, DK-2800 Kongens Lyngby, Denmark
*
Author to whom correspondence should be addressed.
Academic Editor: Holger Barth
Received: 21 October 2016 / Revised: 31 December 2016 / Accepted: 7 January 2017 / Published: 13 Jan 2017
Abstract
A major factor in the resilience of Listeria monocytogenes is the alternative sigma gene B (σB). Type Two Toxin/Antitoxin (TA) systems are also known to take a office in the bacterial stress response upon activation via the ClpP or Lon proteases. Directly upstream of the σB operon in L. monocytogenes is the TA arrangement mazEF, which can cleave mRNA at UACMU sites. In this study, we showed that the mazEF TA locus does not affect the level of persister germination during treatment with antibiotics in lethal doses, but exerts dissimilar effects according to the sub-inhibitory stress added. Growth of a ΔmazEF mutant was enhanced relative to the wildtype in the presence of sub-inhibitory norfloxacin and at 42 °C, merely was decreased when challenged with ampicillin and gentamicin. In contrast to studies in Staphylococcus aureus, we found that the mazEF locus did not bear upon transcription of genes inside the σB operon, merely MazEF effected the expression of the σB-dependent genes opuCA and lmo0880, with a 0.22 and 0.05 fold change, respectively, compared to the wildtype under sub-inhibitory norfloxacin weather condition. How exactly this system operates remains an open question, however, our data indicates it is not analogous to the organisation of S. aureus, suggesting a novel mode of action for MazEF in L. monocytogenes.
ane. Introduction
Listeria monocytogenes is a foodborne pathogenic bacterium, capable of causing major pregnancy complications or severe symptoms, such as septicaemia or meningitis, in immunocompromised persons with upwards of xxx% mortality [1]. In addition to the inhospitable weather condition of the human digestive organisation, Fifty. monocytogenes has evolved to persevere in a several environments; from soil to the intentionally harsh conditions of food processing facilities, where it must cope with exposure to preservatives and disinfectants [two]. One of the most of import factors in the resilience of this organism is the alternative sigma gene B (σB), which under specific weather condition directs RNA polymerase to transcribe >150 stress responsive genes [iii,4] that allow this bacteria to cope with and fifty-fifty abound in NaCl concentrations equally high as 6%, temperatures near 0 °C, and pH as depression as 4.iii [five].
Several other systems are involved in bacterial survival during stress atmospheric condition, including the chromosomally encoded Toxin/Antitoxin (TA) systems, which are most widely known for their role in the formation of persister cells [6]. TA systems are thought to get stochastically activated in a small fraction of a bacterial population, leading to this reversible, fallow-similar cell state that renders the bacterial cell invulnerable to killing past almost classes of antibiotics [6]. I of the most widely studied TA systems is the blazon II TA MazEF. Like all Type II TA systems, the mazEF genes are co-transcribed into 2 proteins, MazE, an unstable antitoxin that binds and inhibits activity of the toxin MazF, an endoribonuclease that cleaves mRNA at specific sites. Under activating conditions of stress in Escherichia coli, mazEF transcription is reduced leading to the degradation of MazE by the ClpP or Lon proteases [7,8], thereby freeing up the toxin to induce stasis via the cleavage of mRNA at ACA sites [ix]. In Staphylococcus aureus, which lacks the Lon protease, MazE is degraded only past the ClpP protease [8] and MazF mRNA cleavage is more specific, occurring at UACAU sites [10]. Persister cells take previously been observed in L. monocytogenes, where the authors speculated that they may result from TA systems [xi]. L. monocytogenes EDGe has ii predicted blazon II TA systems in its chromosome [12], which is relatively few for gratuitous-living prokaryotes [thirteen]. The first is an experimentally verified mazEF TA organization that has been shown to induce dormancy and carve mRNA at UACMU sites upon overexpression [14], while the second is a putative TA organization belonging to the Xre-COG2856 family.
In addition to persister formation, an increasing body of evidence suggests that TA modules also accept a wider regulatory role in stress-related processes, such as the response to nutrient starvation [15] and biofilm formation [16]. Interestingly, the mazEF TA module has been found to exist located direct downstream of the stringent response regulator relA in Gram negative bacteria similar Due east. coli [17] and directly upstream of σB in Gram positive leaner such as Staphylococcus aureus [18], Bacillus subtillis [nineteen], and L. monocytogenes. Reasons for this synteny have and then far been elusive, all the same, Donegan and Cheung [18] have shown that σB of S. aureus motorcar-regulates the σB operon through the mazEF promoter in a stress specific fashion. MazF has also been shown to influence transcript levels of genes within the σB operon [20,21], indicating that it may act as another level of regulation in the σB response.
A second proposed mechanism of action for MazEF presumes that specific genes have evolved to contain unusually high abundances of the MazF cleavage site making them more susceptible to degradation past the toxin under conditions where MazE is degraded. One example is the gene sraP of Southward. aureus, which encodes the pathogenic adhesive factor SraP and contains 43 MazF cleavage sites compared to the xi expected by chance [x]. Furthermore, when the elevation MazF cutting-site rich genes of South. aureus are grouped according to biological office, the pathogenic factor factor group is significantly over-represented [x], whereas the putative MazF target genes in Bacillus subtilis [22] and Staphylococcus equorum [23] are mostly involved in the product of secondary metabolites or metabolism, respectively, suggesting that MazF may regulate unlike specific cellular processes depending on the leaner.
Transcriptomic data from other studies in L. monocytogenes [24,25] show that mazEF (lmo0887-0888), one of only 2 type 2 TA systems predicted in the genome of L. monocytogenes EDGe [12], is likewise co-transcribed in the σB operon. Thus, given the role of the mazEF loci in other organisms, its proximity to the major stress response regulator σB, and the finding that L. monocytogenes forms persister cells via an unknown mechanism [xi], our purpose in this report was to investigate the potential role of this toxin/antitoxin system in the stress response of L. monocytogenes.
2. Results
ii.1. Deletion of the mazEF TA System does not Detectably Modify Persister Formation
Due to an abundance of evidence showing the straight role of TA systems in the formation of persister cells [26,27,28], we initially tested whether or non the deletion of one out of the two predicted TA systems in L. monocytogenes Border would negatively touch on survival when challenged with high concentrations of antibiotics. Upon handling with co-trimoxazole and ampicillin (Figure 1A,B), bacterial counts of each strain did not significantly differ and remained stable throughout the 48 h of exposure. All strains treated with norfloxacin exhibited a biphasic killing curve (Effigy 1C), emblematic of persister cells with an initial steep decline in CFU/mL over the first ten h, followed by a stationary plateau between iv and 3 Log10 CFU/mL for the ΔclpP mutant and between 5.5 and 5.7 Log10 CFU/mL for the remainder. Treatment with gentamicin resulted in a similar biphasic killing curve for all strains (Figure 1D). The only significant departure observed in CFU/mL to EDGe past the terminate of each 48 h handling menstruation was the ΔclpP mutant exposed to norfloxacin (p = 0.001).
ii.2. MazEF Affects Growth under Specific Sub-Inhibitory Growth Conditions
Since no link between the mazEF TA system and persister formation was observed, we hypothesized that information technology could have an agile regulatory function, in which instance information technology may require time to initiate in response to a stress. Therefore, nosotros investigated the furnishings of the factor knockouts on growth under sub-inhibitory stress conditions: specifically, we selected clinically relevant antibiotics, equally well as weather condition known to be alleviated by the σB response. When grown in the absence of treatment (Figure 2A) all strains grew identically, with the exception of the ΔclpP, which was dumb in the exponential growth stage. Relative growth trends in NaCl and bile salts (Effigy 2B,C) were essentially identical to growth in Brain Heart Infusion (BHI) solitary, with the exception that the ΔsigB mutant had a growth advantage when grown in the presence of bile salts. When grown at 42 °C, the ΔmazEF strain had a stark growth reward over the EDGe wildtype strain, and the ΔsigB mutant displayed a moderate growth advantage (Effigy second). In contrast to the killing kinetic experiment, we observed significant differences in growth between the mutants and Border when grown in the presence of sub-inhibitory concentrations of antibiotics (Figure 2E–H). The ΔclpP mutant was more sensitive to all but gentamicin, where its relative growth deficiency to the other strains was abolished (Effigy 2H). For the ΔsigB mutant, we observed slight growth disadvantages relative to EDGe under co-trimoxazole (Figure 2E) and ampicillin (Figure 2F) stress. The ΔmazEF mutant grew more poorly than Border in the presence of co-trimoxazole (Effigy 2E) and ampicillin (Effigy 2F) and was severely inhibited in gentamicin (Figure 2H). However, like to growth at 42 °C, ΔmazEF grew faster and to a higher prison cell density when challenged with norfloxacin (Figure 2G).
In summary, all strains grew identically (except for the ΔclpP mutant) in the absence of stress, however, we observed unique growth differences between the strains depending on the stress added. Nosotros observed similarities betwixt the ΔsigB and ΔmazEF mutants, which grew better at 42 °C and worse in the presence of ampicillin, with the ΔmazEF exhibiting a more extreme difference relative to the wildtype in the two conditions, as well every bit some growth patterns specific for the ΔmazEF mutant, which grew better than the rest of the strains in the presence of norfloxacin, simply worse in gentamicin.
2.iii. MazF Does not Detectably Degrade Putative Target Transcripts In Vivo
Given the established in vitro function of MazF in L. monocytogenes as an endoribonuclease that cleaves mRNAs at UACMU motifs [14], nosotros compared the actual number of MazF cleavage sites in each gene of L. monocytogenes EDGe to the predicted number of MazF cleavage sites, every bit determined by factor length and nucleotide content. This was washed based on the assumption that genes which accept a much higher abundance of MazF cleavage sites, as compared to adventure alone, will be more prone to the ribonucleic activity of MazF. Table ane shows the 10 genes with the highest frequency of MazF cleavage sites, which ranged from functions involved in the electron transport chain (lmo2638), jail cell wall biogenesis and modification (lmo1090 and lmo0835, respectively), and sugar uptake (bvrB) and metabolism (pgi).
To exam if nosotros could discover MazF mediated mRNA cleavage in vivo, we then measured the transcription of three genes with the high frequencies of MazF cut sites: bvrB, lmo2638, and lmo0880 in Border and the ΔmazEF mutant under non-stressed conditions. RT-qPCR showed a slight, just non-significant increase in the number of the three transcripts nowadays in ΔmazEF, as compared to Edge (Effigy three). We also measured the expression of lmo2638 in 50. monocytogenes grown under sub-inhibitory concentrations of NaCl and norfloxacin, merely similarly observed no significant divergence between ΔmazEF and Border (data not shown). The mRNA level of lmo0880, a σB dependent gene, was also measured under sub-inhibitory NaCl and norfloxacin stress (Effigy 4B).
two.four. MazEF Affects Expression of sigB Dependent Genes, but non sigB Operon
In guild to exam if the observed differences in growth were due to a MazEF mediated amending of the σB response, we measured the expression of the σB dependent genes opuCA (glycine betaine-carnitine-choline ABC transporter ATP-bounden protein), lmo0880 (wall associated protein precursor LPXTG motif), and inlA (internalin A) in our strains grown without handling, under osmotic stress (0.5 One thousand NaCl), or with a sub-MIC (Minimum Inhibitory Concentration) (1 µg/mL) of norfloxacin. As expected, expression of each σB dependent gene (Figure four) was decreased in the ΔsigB mutant in each of the tested atmospheric condition with the greatest reductions observed for opuCA and lmo0880 nether stress treatment. Interestingly, in the ΔmazEF mutant, we observed a treatment specific response in opuCA and lmo0880 expression relative to EDGe. We observed no result on opuCA expression in ΔmazEF cultures treated with NaCl, even so, in untreated ΔmazEF cultures, opuCA expression increased when compared to the wildtype (2.549 fold change; p = 0.00006) and decreased when exposed to sub-MIC norfloxacin (0.215 fold modify; p = 0.0004). This trend was reflected for opuCA expression in the ΔclpP mutant, although the only significant deviation (p = 0.0008) was a 0.400 fold change when treated with norfloxacin (Figure 4A). Nosotros also observed a pregnant reduction (0.047 fold alter; p = 0.00001) of lmo0880 expression in the ΔmazEF strain under norfloxacin stress, but no significant difference under no treatment or NaCl handling. These results were once again reflected in the ΔclpP mutant, where lmo0880 expression decreased under norfloxacin pressure level (0.348 fold change; p = 0.00005) and demonstrated similar but non-meaning trends in the other two treatment groups (Figure 4B). Expression of inlA was non significantly different in the ΔclpP or ΔmazEF mutants (Effigy 4C).
These results show a positive influence of σB on opuCA, lmo0880, and inlA expression nether a range of conditions as excepted. On the other mitt, the mazEF gene products appear to repress opuCA expression when cells are unstressed, but, forth with ClpP, activate opuCA and lmo0880 expression when exposed to norfloxacin.
We investigated whether the observed MazEF mediated modification of the σB response was the outcome of transcriptomic regulation of sigB by mazEF, or vice-versa, by quantifying the gene expression of mazF, rsbW (anti-σB factor), and sigB in our strains grown under the same treatment weather. The log2 adapted fold modify of mazF (Figure 5A), rsbW (Effigy 5B), and sigB (Effigy 5C) expression relative to EDGe was less than ii-fold for all of the mutants, signifying that neither σB, ClpP, or MazEF detectably influences the expression of these genes.
Every bit previously mentioned, the promoter of mazEF acts as a stress specific driver of sigB transcription in Due south. aureus [18]. Here we investigated if the mazEF promoter in L. monocytogenes is besides stress responsive by measuring the gene expression of mazF in EDGe cultures receiving either no treatment or exposure to stress (osmotic shock and sub-MIC norfloxacin). Nosotros could not observe a significant difference in expression of mazF across the different treatment groups (Figure six), signifying that the promoter of mazEF in L. monocytogenes is not responsive to the stress atmospheric condition tested.
three. Discussion
Given the established function of the mazEF toxin/antitoxin system in the stress response of several bacterial species [nine,21,26,29,thirty], as well every bit its synteny to the major stress response regulator σB, we hypothesized that this TA system could have an observable function in the response of L. monocytogenes to stress. As chromosomally encoded toxin/antitoxin systems are associated with persister cell formation and tolerance to antibiotics in other bacteria [6,31], nosotros initially conjectured that the mazEF TA organization in L. monocytogenes would serve a similar function, merely this was non the case. Nosotros then hypothesized that the mazEF TA organization could actively regulate the response of L. monocytogenes towards stress, which is supported past the ascertainment that mazEF exerts an effect on the growth of this bacterium under specific sub-inhibitory growth atmospheric condition. TA systems have long been known to induce stasis [32,33], however, this is, to our knowledge, the first written report of a more nuanced role, whereby the TA system exerts different furnishings co-ordinate to the stress (i.e., enhanced growth of the ΔmazEF mutant relative to the wildtype in the presence of norfloxacin and 42 °C, simply decreased growth when challenged with ampicillin and gentamicin) and agrees with the idea that the major office of near TA systems is to serve equally a flexible response of a bacterial prison cell to stress conditions [28,34].
According to the Toxin/Antitoxin Database [12], blazon II TA systems are highly redundant in many bacterial chromosomes, such as E. coli MG1655 strain (16 predicted), Salmonella Typhimurium str. LT2 (18 predicted), and Mycobacterium tuberculosis H37Rv (77 predicted). This back-up means that in some bacteria, multiple TA systems must be removed earlier an event on persister formation can be observed [thirty,35,36]. Thus, it may be possible that the second predicted TA arrangement of 50. monocytogenes EGDe from the Xre-COG2865 family is acting redundantly, however, this TA family unit is essentially uncharacterized and to our knowledge has not been linked to the formation of persister cells. Furthermore, the MazEF TA system appears to exert a ascendant result in certain cases, in that a unmarried deletion of this TA system tin have pregnant effects on the formation of persister cells in E. coli [7] and South. aureus HG003 (nearly isogenic to NCTC8325), fifty-fifty when challenged with a bacteriostatic antibiotic [21]. Conversely, a contempo report in Due south. aureus Newman found that knocking out all known TA systems (including mazEF) had no effect on the level of persisters when challenged with antibiotics, and that persister cells in this Gram positive bacteria are instead the consequence of a stochastic entry into the stationary phase, which the authors speculated could hold true for all Gram positive leaner [37]. Fittingly, the prediction that L. monocytogenes only has two TA systems [12] and our observations that one of these had no detectable consequence on the rate of persister germination is supportive of this theory.
clpP was the only cistron in this study found to touch persister formation. Whether this is due to the bacteria beingness unable to dethrone the antitoxin from the second predicted type II TA organisation of the Xre-COG2856 family (lmo0113-0114), or is the event of an already stressed jail cell due to an inability to clear toxic levels of altered proteins [38], is unknown. However, a reduction in persister cells has recently been shown in a ΔclpP Due south. aureus mutant, where the ΔclpP persister cells were more than sensitive to an antibody from the β-lactam class (oxacillin), just not to a fluoroquinolone (levofloxacin) [39], whereas we observed the opposite with L. monocytogenes. The fact that β-lactams target cell wall biogenesis [40] and fluoroquinolones replicating Dna [41], suggests different roles for ClpP in the germination of persister cells between these two organisms.
The bulk of studies on the MazEF TA organisation take relied on the ectopic overexpression of the toxin to identify its target genes and physiological effects. We, nonetheless, employed a strictly genetic approach, comparing only the presence or absence of the mazEF loci, as we feared that overexpression could accept led to false positives non reflective of this TA system in its natural state. For instance, it seems to us that the conclusion that MazF induces dormancy in S. aureus [20] and 50. monocytogenes [xiv] may exist an artefact of the artificially high levels of toxin in the leaner. While information technology appears to be true for E. coli [7], the observation that no changes in the level of persister cells (a proxy for dormancy) can be observed in ∆mazEF knockouts of South. aureus [37], and as we accept shown hither for 50. monocytogenes, indicates that under natural circumstances the toxin does not accumulate in high plenty concentrations in these Gram positive leaner to induce dormancy. This reasoning was likewise partially why we did not perform a complementation experiment for the ∆mazEF knockout, equally we feared that fifty-fifty a minor difference from the natural levels of MazEF may introduce an artefactual result in the complemented strain. Additionally, we believe that the probability of an off-target mutation occurring in this study was sufficiently depression to justify omitting a complementation experiment, given the method of allelic replacement we used, which employed ii flanking regions with ~450 bp homology to target and delete each gene.
Due to the ribonuclease nature of the MazF toxin in vitro [fourteen], too as its location within the σB operon, we and so hypothesized that mazEF could serve in more than of an active regulatory office and that whatsoever MazF, or potentially related σB mediated effects would require time to manifest. If true, effects of the mazEF deletion would likely exist undetectable when challenged with a sudden exposure to loftier concentrations of antibiotics. To test this, we measured the outcome that each knockout had on growth in sub-inhibitory concentrations of different stressors. Nosotros get-go tested conditions known to stimulate a strong σB response in L. monocytogenes, specifically NaCl [42], bile salts [43], and heat [44]. Interestingly, the only outcome of MazEF on σB associated stress conditions was the enhanced growth of the ΔmazEF mutant at 42 °C, suggesting it plays a part in the thermo-tolerance of L. monocytogenes and is likely the event of heat induced upregulation of mazF [18,45] and/or the ClpP [46] protease, observed in other organisms. Nosotros also observed meliorate growth of the ΔmazEF mutant when grown in sub-inhibitory concentrations of norfloxacin, signifying that under certain conditions, the toxin becomes active and slows the growth of L. monocytogenes, presumably in order to protect the bacteria by reducing the activeness of antibiotic targets [47], which is consequent with studies showing that ectopic induction of artificially high levels of MazF induces stasis in other bacteria [20,48,49].
In contrast, when grown in sub-inhibitory concentrations of ampicillin, the ΔmazEF mutant grew more poorly than the wild type, which correlates with a written report showing that ΔmazEF mutants of Southward. aureus also get more sensitive to β-lactams, where the authors speculate that it may be due to i or more than unknown targets of MazF involved in cell wall synthesis or turnover [21]. Nosotros as well observed an enhanced sensitivity of the ΔmazEF mutant towards gentamicin. This suggests MazEF exerts a protective effect by reducing the membrane potential of the leaner, as aminoglycosides like gentamicin are dependent upon this for their uptake into cells [l]. Given that MazF in L. monocytogenes has been established in vitro as an endoribonuclease that cleaves mRNAs at UACMU motifs [14], besides equally the demonstrated cleavage of the pinnacle putative MazF target genes in Due south. aureus upon mazF overexpression [10], information technology is tempting to explain these growth results through cleavage of the putative MazF target genes, such equally those predicted to be involved in cell wall biogenesis and modification (lmo1090 and lmo0835, respectively) and the electron transport chain (lmo2638, a NADH dehydrogenase). Nonetheless, we were unable to detect a meaning increase in the transcript levels of the three putative MazF target genes we tested in the ∆mazEF mutant every bit compared to the wildtype. The discrepancy between the MazF mediated mRNA cleavage in vitro and a lack of cleavage in vivo has besides been observed in Southward. aureus and is thought to be the result of protective RNA bounden proteins [20]. Furthermore, the observation that lmo0880, which is both a σB dependent and putative MazF target cistron, is significantly decreased under norfloxacin stress in the ∆mazEF mutant contrasts with the idea of MazF mediated cleavage of transcripts containing high frequencies of the UACMU motif in L. monocytogenes.
We also investigated the possibility that MazEF affects growth nether stress by modifying the σB response, which is supported past our observation that MazEF activates the expression of the σB dependent genes opuCA and lmo0880 nether norfloxacin stress. However, we observed that inlA expression was non affected by the absence of MazEF, but speculate that the effect may exist diluted for this gene due to its co-regulation by the global virulence regulator PrfA [51]. In a recent written report on the transcriptomic response to sub-lethal antibiotics [25], L. monocytogenes was institute to significantly down-regulate the expression of certain σB dependent genes, including opuCA and lmo0880, in response to all tested antibiotics. Our observations advise that MazEF tempers this antibiotic induced downregulation of these genes, thereby interim as a negative feedback mechanism to fine tune this response, which is consistent with the hypothesis that TA systems act as a quality command mechanism for gene expression under stress [28].
In an attempt to characterize the directly mechanism through which MazEF influences the expression of opuCA, lmo0880, and potentially other σB-dependent genes, we get-go examined whether or not MazEF directly modifies the expression of genes within the σB operon, equally has been observed in S. aureus for sigB [20] and rsbW [21], nevertheless, we were unable to observe whatever event on transcript levels of either cistron in the presence or absence of mazEF. We then tested some other link observed in S. aureus between mazEF and the σB operon discovered by Donegan and Cheung [xviii] who, using an experimental setup similar to ours, showed that transcripts of mazF in South. aureus were significantly elevated in their ΔsigB strain, and that the complete 3.7 kb σB operon transcript (mazEF-sigB) was present only when exposed to certain conditions, such equally heat stupor or sub-inhibitory concentrations of erythromycin and tetracycline, but non to vancomycin. This, in effect, means that the promoter of mazEF in S. aureus acts as a stress specific promoter for the σB operon that is machine-repressed by σB, thereby allowing the bacterium to fine tune its response to item stimuli. This would explain the provisional effect of mazEF on opuCA and lmo0880 expression, however, we observed no significant furnishings of either sub-inhibitory NaCl or norfloxacin on the expression of mazF, which would be expected if the promoter of mazEF in L. monocytogenes were acting in an analogous style. Furthermore, σB had no effect on the expression of mazF in our study, or mazE equally reported past other transcriptomic studies on ΔsigB mutants of Fifty. monocytogenes [52,53], indicating that information technology does not act in the automobile-regulatory manner observed in South. aureus. Our results advise that mazEF modulates the σB response via some novel mechanism in L. monocytogenes, peradventure past cleaving the mRNA of the poly peptide or proteins responsible for affecting the antibiotic induced downregulation of these genes.
four. Conclusions
At the physiological level, mazEF appears not to stop growth, equally evidenced past its zero effect on dormant persister cell formation, but rather modulate growth in response to specific stress factors. At the transcriptional level, MazEF modifies the expression of certain σB dependent genes similar opuCA and lmo0880 in a stress specific manner. Interestingly, with the exception of heat, mazEF seems immaterial with respect to weather condition typically associated with the σB response (i.east., NaCl and bile), suggesting that information technology may piece of work past co-opting and redirecting the pre-existing σB stress response pathway to cope with additional stressors like antibiotics. How exactly this system operates remains an open up question, however, we have ruled out similar models based on inquiry in S. aureus, suggesting a novel mode of MazEF action in L. monocytogenes.
5. Materials and Methods
5.i. Bacterial Strains and Growth Conditions
L. monocytogenes EGDe wild-type was used in the written report (Table 2). Bacterial stock cultures were stored at −fourscore °C and inoculated on Brain Heart Infusion (BHI; Oxoid CM 1135) agar and grown at 37 °C overnight. An overnight culture was obtained by inoculating one colony in x mL BHI broth and incubating aerobically at 37 °C with shaking (250 rpm).
5.2. Antibiotic Preparation
Fresh antibiotic solutions were prepared for each experiment: ampicillin (dissolved in sterile MilliQ h2o; Sigma-Aldrich A9518, St. Louis, MO, Us), gentamicin (dissolved in sterile MilliQ h2o; Sigma-Aldrich G3632), norfloxacin (dissolved in sterile MilliQ with 1% glacial acetic acid; Fluka N9890), and co-trimoxazole, which is comprised of one office trimethoprim (dissolved in sterile MilliQ with i% glacial acerb acid; Sigma-Aldrich 92131) and five parts sulphamethoxazole (dissolved in acetone; Fluka S7507).
v.3. Constructions of Mutants
In-frame deletion mutants were synthetic in the sigB, clpP, mazEF genes using the gene splicing past overlap extension (gene SOEing) method [55]. Primers (Tabular array 3) were constructed using the published sequence of 50. monocytogenes EGD-e (31). Chromosomal DNA from EGDe was used as the PCR template for the SOEing amplicons, which were subsequently cloned into the pAUL-A vector [54]. Standard protocols were used for plasmid extractions, brake enzyme digests, and Dna ligations [56]. Plasmids harboring the SOEing fragments were isolated and verified past sequencing. Generation of the deletion mutants was performed equally described by Guzman et al. [57], with the exception that electrocompetent cells were prepared every bit described in Monk et al. [58]. Putative mutants were verified by sequencing at GATC (Koln, Deutschland). The morphology of each strain was determined to be identical using an Olympus BH2 microscope at 1000X magnification.
5.4. Killing Kinetics
Standard killing kinetic experiments were performed every bit previously described [11], with the exception that the second regrowth and killing footstep was omitted. In brief, an overnight culture from each strain was diluted 10six-fold and grown for 16 h at 37 °C at 250 rpm to obtain an early stationary phase civilisation. This xvi h civilization was diluted to OD600 = 0.4 in 2 mL BHI broth and treated with roughly 30 times the Minimum Inhibitory Concentration (MIC) of four bactericidal antibiotics: norfloxacin (100 µg/mL), co-trimoxazole (x µg/mL), ampicillin (3 µg/mL), and gentamicin (x µg/mL). Cultures were incubated at 37 °C and 250 rpm during antibiotic treatment. Bacterial counts were determined merely earlier treatment and at subsequent time points by plate counting on BHI agar. The experiment was performed with three biological replicates.
v.5. Growth Kinetics
Overnight cultures of each strain were adapted to OD600 = 0.ii and diluted by 10four-fold, and so the Minimum Inhibitory Concentration (MIC) for each treatment (antibiotics, 42 °C, NaCl, and Bile acid) was identified using a standard two-fold dilution serial in 96-well plates using ii biological replicates for each strain. Plates were incubated at 37 °C for 24 h in the SpectraMax i3 (Molecular Devices) Multi-Mode microplate reader ready to measure OD600 every 60 minutes. In one case the MIC for each treatment was identified, the same procedure higher up was carried out using two biological replicates, with the exception that the highest concentration started with the MIC and was serially diluted by 10%. Sub-inhibitory concentrations were chosen on the basis of being closest to the MIC, while still allowing for exponential and stationary growth curves of Border, ΔsigB, and ΔmazEF. Antibiotics were prepared as described above, and fresh solutions of NaCl (Merck) and Bile Salts (Cholic acrid sodium salt 50% and Deoxycholic acid sodium salt 50%; Fluka, 48305-50G-F) were dissolved in sterile MilliQ water.
five.6. Sample Collection and RNA Isolation
To obtain balanced growth, overnight cultures of each strain were diluted 109-fold in ten mL BHI broth and incubated aerobically at 37 °C with shaking (250 rpm) for 16 h, then subsequently diluted to OD600 0.01 in 50 mL pre-warmed BHI in a 500 mL flask. Upon reaching OD600 0.1, each culture was split up into three 100 mL flasks, past taking vii mL of civilisation into 23 mL pre-warmed BHI, and farther incubated to OD600 0.1. Flasks received either 3 mL MQ water for the no treatment control, or three mL MQ water containing NaCl or norfloxacin, yielding a final concentration of 0.5 M or ane µg/mL, respectively. Each culture was incubated to OD600 0.forty, then 5 mL culture was immediately added to ten mL RNA-protect Leaner Reagent (Qiagen, 76506, Hilden, Federal republic of germany) and after treated according to the manufacturer's instructions. Two independent biological replicates for each strain were carried out.
Cell lysis for downstream RNA purification was carried out by first resuspending each isolated pellet in 500 µL Buffer RLT (Qiagen, 74104) with 5 µL β-meracaptoethanol. 3 hundred milligrams of acid washed glass beads were added to the cell suspension, and lysed using the FastPrep FP120 Cell Disruptor. Samples were centrifuged at 4500× chiliad for 2 min at 4 °C, then 500 µL of the cell lysis supernatant was mixed with 250 µL 96% ethanol and farther processed using the RNeasy mini kit (Qiagen, 74104) according to manufacturer's protocol "Purification of total RNA from bacteria", using the additional on column DNase treatment step. RNA quality and quantity were measured using the Agilent 2100 Bioanalyser with a RNA 6000 Nano bit and the DeNovix DS-11+ spectrophotometer, respectively. All RIN values were above 9, with the exception of one replicate of the ΔclpP under non-stressed conditions, which had a RIN value of seven.6.
five.7. Measuring Relative Gene Expression via Quantitative Reverse Transcription PCR (RT-qPCR)
Prior to cDNA synthesis, 2 µg of RNA from each sample was treated with DNase I (Invitrogen, 18068-015, Carlsbad, CA, Usa) and dissever into opposite transcriptase positive (RT+) and negative (RT−) tubes. cDNA synthesis was carried out for RT+ samples using SuperScript Three Reverse Transcriptase (Invitrogen, 18080-044) and random primers (Invitrogen, 48190-011) according to the manufacturer's protocol for First-Strand cDNA Synthesis. RT− samples were treated identically, with the exception that MQ water was substituted for contrary transcriptase. Prior to qPCR templet cDNA was diluted 4× in MQ h2o except when investigating 16S, which was diluted 1000×.
RT-qPCR was performed using SYBR Green PCR Master Mix (Applied Biosystems, Foster City, CA, Usa) on the Mx3000P (Stratagene, San Diego, CA, U.s.) using the following plan: one cycle at 95 °C for x min, followed by 40 cycles at 95 °C for 30 due south and 60 °C for 1 min followed past a dissociation curve. Three technical replicates for each RT+ cDNA sample and two technical replicates for RT− cDNA sample were performed for each reaction, along with negative template control using MQ water. Primers (Table 3) were designed using Primer3 [59] and standard curves for each primer set were done using a dilution series of Border genomic Deoxyribonucleic acid. 16S gene expression has previously been demonstrated to be stable under stress weather condition [threescore], and was shown in this experiment to be like between each strain and condition tested using an ANOVA test. 16S factor expression was measured and used to normalize expression levels of each gene measured, which were further normalized to Border expression levels using the 2−ΔΔCt method [61]. When comparing the expression of the mazF cistron alone under different atmospheric condition, copy numbers for mazF and 16S rRNA gene transcripts were calculated and expressed every bit target (mazF)/housekeeping (16S rRNA factor).
5.8. Bioinformatic Assay of MazF Cut Site Abundance in Edge Genome
In lodge to predict the relative abundance of the UACMU motif per gene, as compared to gamble, the coding sequences (CDS) of L. monocytogenes EDGe (accession No. NC_003210.1) were run through a custom python script based on the formula described in Zhu et al., 2009 [10]:
Briefly, the probability (p) of the UACMU motif occurring in a CDS was calculated using the length (Fifty) and base composition, which equals (%U's)ii(%A's)2(%C's) + (%U's)2(%A's) (%C's)2. Thus, the expected number of motifs in each CDS equals p(L-4). If K is the actual number of motifs found in a CDS, so P equals the probability of a CDS containing K or fewer motifs. Source code has been uploaded to GitHub [62].
5.9. Statistical Analysis
Bacterial counts and OD600 measurements from each biological replicate were logten transformed, whereas ΔΔCt values from RT-qPCR were logii transformed, prior to statistical assay using the macro, Analysis ToolPak, in Microsoft Excel. The F-Test was used to test for equal Variances of the sample populations and Students t-test with equal or unequal variance was used when advisable, with a significance level of p < 0.05.
Acknowledgments
This work was supported by a Ph.D. grant from the Department of Biotechnology and Biomedicine, Technical University of Denmark. We would like to thank members of the Gram Lab for expert technical assistance and good discussions, peculiarly Yin Ng for her work on the killing kinetics. Nosotros would likewise like to thank Asker Brejnrod from the University of Copenhagen for his assist with Python.
Author Contributions
Conceived and designed the experiments: T.D.C., I.T., L.M. and G.M.Grand. Performed the experiments: T.D.C. and I.T. Analyzed the information: T.D.C., L.G. and G.M.1000. Wrote the manuscript: T.D.C., L.K. and Thou.Grand.K.
Conflicts of Involvement
The authors declare no conflict of involvement.
References
- Drevets, D.A.; Bronze, M.Due south. Listeria monocytogenes: Epidemiology, human illness, and mechanisms of brain invasion. FEMS Immunol. Med. Microbiol. 2008, 53, 151–165. [Google Scholar] [CrossRef] [PubMed]
- Freitag, Northward.Due east.; Port, G.C.; Miner, M.D. Listeria monocytogenes—From saprophyte to intracellular pathogen. Nat. Rev. Microbiol. 2009, 7, 623–628. [Google Scholar] [CrossRef] [PubMed]
- Becker, L.A.; Cetin, M.S.; Hutkins, R.W.; Benson, A.K. Identification of the gene encoding the alternative sigma cistron sigmaB from Listeria monocytogenes and its office in osmotolerance. J. Bacteriol. 1998, 180, 4547–4554. [Google Scholar] [PubMed]
- Wiedmann, Yard.; Arvik, T.J.; Hurley, R.J.; Boor, K.J. General stress transcription factor sigmaB and its office in acrid tolerance and virulence of Listeria monocytogenes. J. Bacteriol. 1998, 180, 3650–3656. [Google Scholar] [PubMed]
- Tienungoon, S.; Ratkowsky, D.A.; McMeekin, T.A.; Ross, T. Growth limits of Listeria monocytogenes every bit a function of temperature, pH, NaCl, and lactic acrid. Appl. Environ. Microbiol. 2000, 66, 4979–4987. [Google Scholar] [CrossRef] [PubMed]
- Maisonneuve, Due east.; Gerdes, K. Molecular mechanisms underlying bacterial persisters. Cell 2014, 157, 539–548. [Google Scholar] [CrossRef] [PubMed]
- Tripathi, A.; Dewan, P.C.; Siddique, S.A.; Varadarajan, R. MazF-induced growth inhibition and persister generation in Escherichia coli. J. Biol. Chem. 2014, 289, 4191–4205. [Google Scholar] [CrossRef] [PubMed]
- Donegan, North.P.; Thompson, E.T.; Fu, Z.; Cheung, A.L. Proteolytic regulation of toxin-antitoxin systems by ClpPC in Staphylococcus aureus. J. Bacteriol. 2010, 192, 1416–1422. [Google Scholar] [CrossRef] [PubMed]
- Christensen, S.1000.; Pedersen, K.; Hansen, F.G.; Gerdes, K. Toxin-antitoxin loci every bit stress-response-elements: ChpAK/MazF and ChpBK cleave translated RNAs and are counteracted by tmRNA. J. Mol. Biol. 2003, 332, 809–819. [Google Scholar] [CrossRef]
- Zhu, Fifty.; Inoue, 1000.; Yoshizumi, S.; Kobayashi, H.; Zhang, Y.; Ouyang, M.; Kato, F.; Sugai, M.; Inouye, K. Staphylococcus aureus MazF specifically cleaves a pentad sequence, UACAU, which is unusually abundant in the mRNA for pathogenic adhesive factor SraP. J. Bacteriol. 2009, 191, 3248–3255. [Google Scholar] [CrossRef] [PubMed]
- Knudsen, G.Yard.; Ng, Y.; Gram, 50. Survival of bactericidal antibiotic treatment by a persister subpopulation of Listeria monocytogenes. Appl. Environ. Microbiol. 2013, 79, 7390–7397. [Google Scholar] [CrossRef] [PubMed]
- Shao, Y.; Harrison, E.M.; Bi, D.; Tai, C.; He, X.; Ou, H.-Y.; Rajakumar, 1000.; Deng, Z. TADB: A web-based resource for Type two toxin-antitoxin loci in leaner and archaea. Nucleic Acids Res. 2011, 39, D606–D611. [Google Scholar] [CrossRef] [PubMed]
- Pandey, D.P.; Gerdes, Yard. Toxin-antitoxin loci are highly arable in free-living but lost from host-associated prokaryotes. Nucleic Acids Res. 2005, 33, 966–976. [Google Scholar] [CrossRef] [PubMed]
- Schmitter, S. Part of a Toxin-Antitoxin System in Fifty. monocytogenes. Doctoral and Habilitation Thesis, Eidgenossische Technische Hochschule ETH Zurich, Zürich, Switzerland, 2014. [Google Scholar]
- Christensen, South.Grand.; Gerdes, K. Delayed-relaxed response explained by hyperactivation of RelE. Mol. Microbiol. 2004, 53, 587–597. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez Barrios, A.F.; Zuo, R.; Hashimoto, Y.; Yang, L.; Bentley, W.E.; Wood, T.K. Autoinducer 2 Controls Biofilm Germination in Escherichia coli through a Novel Motility Quorum-Sensing Regulator (MqsR, B3022). J. Bacteriol. 2006, 188, 305–316. [Google Scholar] [CrossRef] [PubMed]
- Metzger, South.; Dror, I.B.; Aizenman, E.; Schreiber, G.; Toone, M.; Friesen, J.D.; Cashel, M.; Glaser, Yard. The nucleotide sequence and characterization of the relA gene of Escherichia coli. J. Biol. Chem. 1988, 263, 15699–15704. [Google Scholar] [PubMed]
- Donegan, N.P.; Cheung, A.L. Regulation of the mazEF toxin-antitoxin module in Staphylococcus aureus and its impact on sigB expression. J. Bacteriol. 2009, 191, 2795–2805. [Google Scholar] [CrossRef] [PubMed]
- Pellegrini, O.; Mathy, N.; Gogos, A.; Shapiro, 50.; Condon, C. The Bacillus subtilis ydcDE operon encodes an endoribonuclease of the MazF/PemK family unit and its inhibitor. Mol. Microbiol. 2005, 56, 1139–1148. [Google Scholar] [CrossRef] [PubMed]
- Fu, Z.; Tamber, S.; Memmi, G.; Donegan, N.P.; Cheung, A.L. Overexpression of MazFsa in Staphylococcus aureus induces bacteriostasis by selectively targeting mRNAs for cleavage. J. Bacteriol. 2009, 191, 2051–2059. [Google Scholar] [CrossRef] [PubMed]
- Schuster, C.F.; Mechler, L.; Nolle, N.; Krismer, B.; Zelder, M.-East.; Gotz, F.; Bertram, R. The MazEF Toxin-Antitoxin System Alters the β-Lactam Susceptibility of Staphylococcus aureus. PLoS One 2015, x. [Google Scholar] [CrossRef]
- Park, J.H.; Yamaguchi, Y.; Inouye, Thou. Bacillus subtilis MazF-bs (EndoA) is a UACAU-specific mRNA interferase. FEBS Lett. 2011, 585, 2526–2532. [Google Scholar] [CrossRef] [PubMed]
- Schuster, C.F.; Park, J.H.; Prax, M.; Herbig, A.; Nieselt, K.; Rosenstein, R.; Inouye, Thousand.; Bertrama, R. Characterization of a MazEF toxin-antitoxin homologue from Staphylococcus equorum. J. Bacteriol. 2013, 195, 115–125. [Google Scholar] [CrossRef] [PubMed]
- Wurtzel, O.; Sesto, N.; Mellin, J.R.; Karunker, I.; Edelheit, Southward.; Becavin, C.; Archambaud, C.; Cossart, P.; Sorek, R. Comparative transcriptomics of pathogenic and not-pathogenic Listeria species. Mol. Syst. Biol. 2012, viii. [Google Scholar] [CrossRef] [PubMed]
- Knudsen, G.M.; Fromberg, A.; Ng, Y.; Gram, 50. Sublethal Concentrations of Antibiotics Cause Shift to Anaerobic Metabolism in Listeria monocytogenes and Induce Phenotypes Linked to Antibiotic Tolerance. Front. Microbiol. 2016, vii, 1091. [Google Scholar] [CrossRef] [PubMed]
- Merfa, M.5.; Niza, B.; Takita, M.A.; De Souza, A.A. The MqsRA Toxin-Antitoxin Organisation from Xylella fastidiosa Plays a Key Office in Bacterial Fitness, Pathogenicity, and Persister Cell Formation. Front. Microbiol. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Lobato-Marquez, D.; Moreno-Cordoba, I.; Figueroa, V.; Diaz-Orejas, R.; Garcia-del Portillo, F. Singled-out blazon I and type II toxin-antidote modules control Salmonella lifestyle inside eukaryotic cells. Sci. Rep. 2015, 5. [Google Scholar] [CrossRef] [PubMed]
- Gerdes, Thou.; Christensen, South.K.; Lobner-Olesen, A. Prokaryotic toxin-antitoxin stress response loci. Nat. Rev. Microbiol. 2005, 3, 371–382. [Google Scholar] [CrossRef] [PubMed]
- Lemos, J.A.C.; Chocolate-brown, T.A.; Abranches, J.; Burne, R.A. Characteristics of Streptococcus mutans strains defective the MazEF and RelBE toxin-antitoxin modules. FEMS Microbiol. Lett. 2005, 253, 251–257. [Google Scholar] [CrossRef] [PubMed]
- Silva-Herzog, E.; McDonald, E.G.; Crooks, A.L.; Detweiler, C.S. Physiologic Stresses Reveal a Salmonella Persister State and TA Family Toxins Modulate Tolerance to These Stresses. PLoS 1 2015, ten. [Google Scholar] [CrossRef] [PubMed]
- Page, R.; Peti, W. Toxin-antitoxin systems in bacterial growth arrest and persistence. Nat. Chem. Biol. 2016, 12, 208–214. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, G.; Christensen, S.K.; Gerdes, K. Rapid induction and reversal of a bacteriostatic condition by controlled expression of toxins and antitoxins. Mol. Microbiol. 2002, 45, 501–510. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, M.; Zhang, J.; Liu, One thousand.; Woychik, Northward.A.; Inouye, One thousand. Unmarried protein production in living cells facilitated past an mRNA interferase. Mol. Cell 2005, 18, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Bukowski, K.; Rojowska, A.; Wladyka, B. Prokaryotic toxin-antitoxin systems--the function in bacterial physiology and application in molecular biological science. Acta Biochim. Pol. 2011, 58, 1–ix. [Google Scholar] [PubMed]
- Maisonneuve, E.; Gerdes, Yard. Supporting Information: Bacterial persistence by RNA endonucleases. Proc. Natl. Acad. Sci. USA 2011, 108, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, P.; Arora, G.; Singh, M.; Kidwai, S.; Narayan, O.P.; Singh, R. MazF ribonucleases promote Mycobacterium tuberculosis drug tolerance and virulence in guinea pigs. Nat. Commun. 2015, vi. [Google Scholar] [CrossRef] [PubMed]
- Conlon, B.P.; Rowe, Southward.E.; Gandt, A.B.; Nuxoll, A.S.; Donegan, N.P.; Zalis, E.A.; Clair, G.; Adkins, J.N.; Cheung, A.Fifty.; Lewis, K. Persister germination in Staphylococcus aureus is associated with ATP depletion. Nat. Microbiol. 2016, 1. [Google Scholar] [CrossRef] [PubMed]
- Gaillot, O.; Pellegrini, E.; Bregenholt, Southward.; Nair, S.; Berche, P. The ClpP serine protease is essential for the intracellular parasitism and virulence of Listeria monocytogenes. Mol. Microbiol. 2000, 35, 1286–1294. [Google Scholar] [CrossRef] [PubMed]
- Springer, One thousand.T.; Singh, V.K.; Cheung, A.L.; Donegan, N.P.; Chamberlain, N.R. Issue of clpP and clpC deletion on persister cell number in Staphylococcus aureus. J. Med. Microbiol. 2016, 65, 848–857. [Google Scholar] [CrossRef] [PubMed]
- Page, M. The Chemical science of β-Lactams; Folio, One thousand.I., Ed.; Springer: Dordrecht, The Netherlands, 1992. [Google Scholar]
- Blondeau, J.M. Fluoroquinolones: Mechanism of action, nomenclature, and evolution of resistance. Surv. Ophthalmol. 2004, 49 (Suppl. 2), S73–S78. [Google Scholar] [CrossRef] [PubMed]
- Kazmierczak, G.J.; Wiedmann, M.; Boor, Grand.J. Contributions of Listeria monocytogenes SigmaB and PrfA to expression of virulence and stress response genes during extra- and intracellular growth. Microbiology 2006, 152, 1827–1838. [Google Scholar] [CrossRef] [PubMed]
- Sue, D.; Churl, Thou.J.; Wiedmann, M. Sigma(B)-dependent expression patterns of compatible solute transporter genes opuCA and lmo1421 and the conjugated bile table salt hydrolase cistron bsh in Listeria monocytogenes. Microbiology 2003, 149, 3247–3256. [Google Scholar] [CrossRef] [PubMed]
- McGann, P.; Ivanek, R.; Wiedmann, Grand.; Boor, K.J. Temperature-dependent expression of Listeria monocytogenes internalin and internalin-similar genes suggests functional diversity of these proteins amongst the listeriae. Appl. Environ. Microbiol. 2007, 73, 2806–2814. [Google Scholar] [CrossRef] [PubMed]
- Muthuramalingam, M.; White, J.C.; Bourne, C.R. Toxin-Antitoxin Modules Are Pliable Switches Activated by Multiple Protease Pathways. Toxins 2016, eight, 214. [Google Scholar] [CrossRef] [PubMed]
- Gerth, U.; Kruger, E.; Derre, I.; Msadek, T.; Hecker, M. Stress consecration of the Bacillus subtilis clpP gene encoding a homologue of the proteolytic component of the Clp protease and the interest of ClpP and ClpX in stress tolerance. Mol. Microbiol. 1998, 28, 787–802. [Google Scholar] [CrossRef] [PubMed]
- Prax, One thousand.; Bertram, R. Metabolic aspects of bacterial persisters. Front. Cell. Infect. Microbiol. 2014, iv, ane–6. [Google Scholar] [CrossRef] [PubMed]
- Syed, Thousand.A.; Koyanagi, S.; Sharma, E.; Jobin, Grand.C.; Yakunin, A.F.; Levesque, C.M. The chromosomal mazEF locus of Streptococcus mutans encodes a functional blazon II toxin-antitoxin addiction system. J. Bacteriol. 2011, 193, 1122–1130. [Google Scholar] [CrossRef] [PubMed]
- Mok, W.W.K.; Park, J.O.; Rabinowitz, J.D.; Brynildsen, Grand.P. RNA Futile Cycling in Model Persisters Derived from MazF Aggregating. MBio 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Mingeot-Leclercq, M.-P.; Glupczynski, Y.; Tulkens, P.M. Aminoglycosides: Activity and Resistance. Antimicrob. Agents Chemother. 1999, 43, 727–737. [Google Scholar] [PubMed]
- Scortti, K.; Monzo, H.J.; Lacharme-Lora, L.; Lewis, D.A.; Vazquez-Boland, J.A. The PrfA virulence regulon. Microbes Infect. 2007, 9, 1196–1207. [Google Scholar] [CrossRef] [PubMed]
- Oliver, H.F.; Orsi, R.H.; Ponnala, Fifty.; Keich, U.; Wang, W.; Sun, Q.; Cartinhour, S.Westward.; Filiatrault, M.J.; Wiedmann, M.; Boor, Chiliad.J. Deep RNA sequencing of L. monocytogenes reveals overlapping and extensive stationary phase and sigma B-dependent transcriptomes, including multiple highly transcribed noncoding RNAs. BMC Genom. 2009, ten. [Google Scholar] [CrossRef] [PubMed]
- Hain, T.; Hossain, H.; Chatterjee, S.Due south.; Machata, Southward.; Volk, U.; Wagner, Due south.; Brors, B.; Haas, South.; Kuenne, C.T.; Billion, A.; et al. Temporal transcriptomic analysis of the Listeria monocytogenes EGD-due east sigmaB regulon. BMC Microbiol. 2008, eight. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, T.; Leimeister-Wachter, M.; Domann, E.; Hartl, Chiliad.; Goebel, Westward.; Nichterlein, T.; Notermans, South. Coordinate regulation of virulence genes in Listeria monocytogenes requires the product of the prfA gene. J. Bacteriol. 1992, 174, 568–574. [Google Scholar] [CrossRef] [PubMed]
- Horton, R.Grand.; Cai, Z.; Ho, S.N.; Pease, L.R. Factor splicing by overlap extension: Tailor-made genes using the polymerase concatenation reaction. Biotechniques 2013, 54, 528–535. [Google Scholar] [CrossRef]
- Sambrook, J.; Russell, D.W. Molecular Cloning: A Laboratory Transmission; Cold Jump Harbor Laboratory Press: New York, NY, United states of america, 2001; p. 999. [Google Scholar]
- Guzman, C.A.; Rohde, M.; Chakraborty, T.; Domann, E.; Hudel, G.; Wehland, J.; Timmis, K.N. Interaction of Listeria monocytogenes with mouse dendritic cells. Infect. Immun. 1995, 63, 3665–3673. [Google Scholar] [PubMed]
- Monk, I.R.; Gahan, C.G.M.; Loma, C. Tools for functional postgenomic analysis of Listeria monocytogenes. Appl. Environ. Microbiol. 2008, 74, 3921–3934. [Google Scholar] [CrossRef] [PubMed]
- Primer3. Available online: http://frodo.wi.mit.edu/primer3/input.htm (accessed on xiv May 2016).
- Tasara, T.; Stephan, R. Evaluation of housekeeping genes in Listeria monocytogenes as potential internal control references for normalizing mRNA expression levels in stress accommodation models using real-time PCR. FEMS Microbiol. Lett. 2007, 269, 265–272. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of Relative Gene Expression Data Using Existent-Time Quantitative PCR and the 2−ΔΔCt Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef] [PubMed]
- GitHub. Available online: https://github.com/thocu/MazF_Targets.git (accessed on 7 Oct 2016).
Figure 1. Killing kinetics of EDGe (○), ΔsigB (■), ΔclpP (♦), and ΔmazEF (□) exposed to approximately 30× Minimum Inhibitory Concentrations (MIC) for 72 h under shaking at 250 rpm with (A) co-trimoxazole (x µg/mL); (B) ampicillin (3 µg/mL); (C) norfloxacin (100 µg/mL); and (D) gentamicin (10 µg/mL). Error bars represent standard deviation of the mean for three biological replicates. * p < 0.001.
Effigy i. Killing kinetics of EDGe (○), ΔsigB (■), ΔclpP (♦), and ΔmazEF (□) exposed to approximately 30× Minimum Inhibitory Concentrations (MIC) for 72 h under shaking at 250 rpm with (A) co-trimoxazole (10 µg/mL); (B) ampicillin (3 µg/mL); (C) norfloxacin (100 µg/mL); and (D) gentamicin (x µg/mL). Error confined represent standard deviation of the mean for iii biological replicates. * p < 0.001.

Figure two. Growth curves for Border (○), ΔsigB (■), ΔclpP (♦), and ΔmazEF (□) exposed to sub-inhibitory stress. OD600 was measured in 96-well plates over 24 h for strains receiving (A) no handling; (B) 0.625 M NaCl; (C) 0.01% bile salts; (D) 42 °C; (Eastward) 0.25 µg/mL co-trimoxazole (SXT); (F) 0.07 µg/mL ampicillin (AMP); (G) 1.v µg/mL norfloxacin (NOR); and (H) 0.10 µg/mL gentamicin (GEN). Fault bars represent standard deviation of the hateful for ii biological replicates.
Figure 2. Growth curves for Edge (○), ΔsigB (■), ΔclpP (♦), and ΔmazEF (□) exposed to sub-inhibitory stress. OD600 was measured in 96-well plates over 24 h for strains receiving (A) no treatment; (B) 0.625 M NaCl; (C) 0.01% bile salts; (D) 42 °C; (Due east) 0.25 µg/mL co-trimoxazole (SXT); (F) 0.07 µg/mL ampicillin (AMP); (G) 1.five µg/mL norfloxacin (NOR); and (H) 0.10 µg/mL gentamicin (GEN). Error bars represent standard deviation of the mean for two biological replicates.

Figure 3. Quantification of putative MazF target mRNA via RT-qPCR. Fold-change in Logtwo scale of the ΔmazEF mutant relative to EDGe under not-stressed conditions is shown for lmo2638, lmo0880, and bvrB, with one, 2, and 0 MazF cleavage motifs inside the amplicon, respectively. Error confined represent standard departure of the hateful for two biological replicates.
Figure iii. Quantification of putative MazF target mRNA via RT-qPCR. Fold-alter in Logtwo calibration of the ΔmazEF mutant relative to EDGe under not-stressed conditions is shown for lmo2638, lmo0880, and bvrB, with 1, 2, and 0 MazF cleavage motifs within the amplicon, respectively. Fault confined stand for standard deviation of the hateful for two biological replicates.
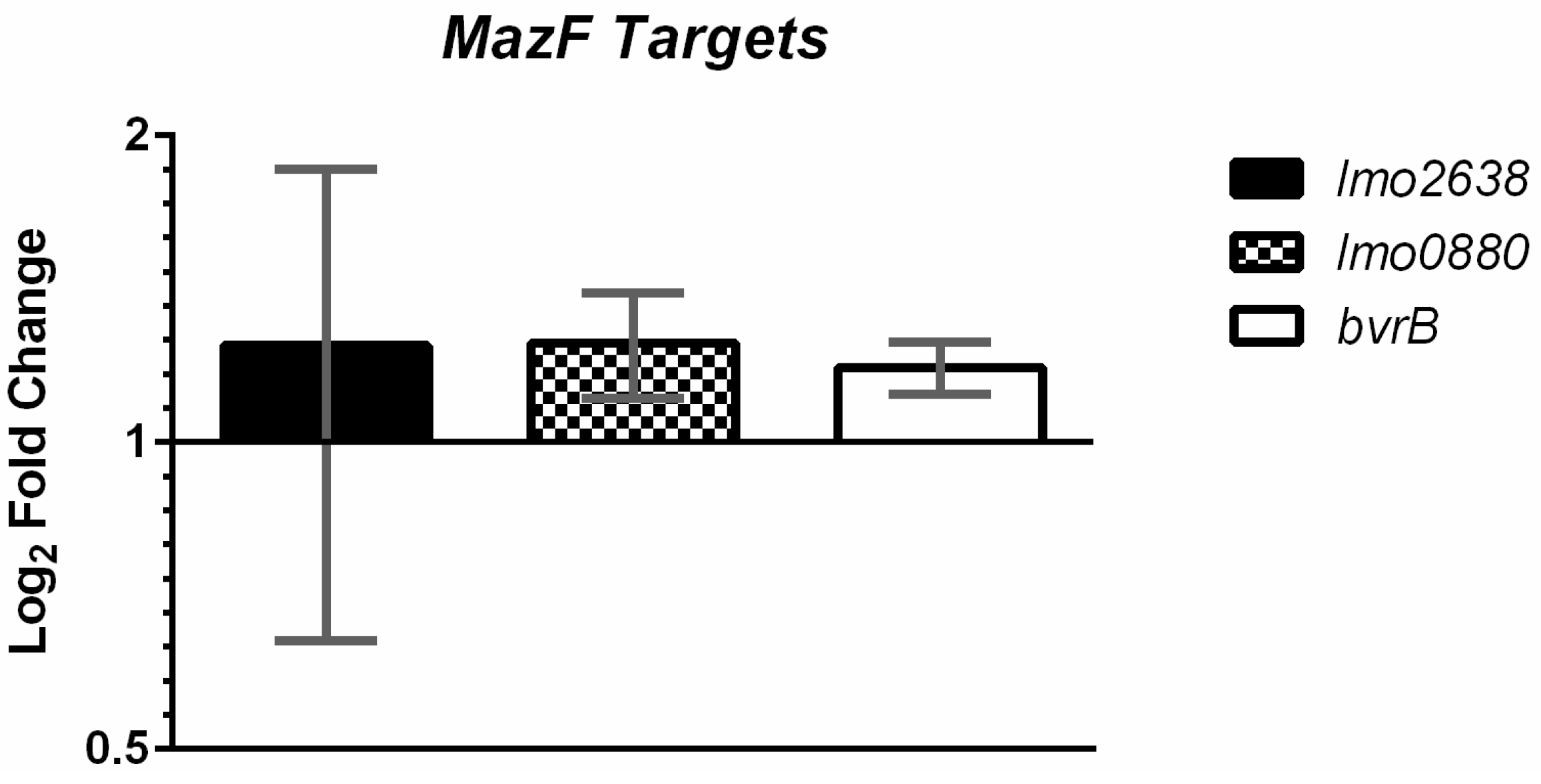
Figure iv. Quantification of σB-dependent gene expression via RT-qPCR. Fold-change in Log2 scale of each gene relative to Border is shown for the (A) opuCA; (B) lmo0880; and (C) inlA genes. Each strain was grown at 37 °C with 200 rpm shaking and received either no treatment (MQ), osmotic stress (0.v M NaCl), or sub-MIC (1 µg/mL) norfloxacin. Ct values were normalized to the 16S rRNA gene (ΔCt), and and then compared to EDGe expression (ΔΔCt), and the average was taken from two independent experiments. Significance was determined using a t-exam on the logii transformed ii − ΔΔCt values. * p < 0.05, ** p < 0.0005, *** p < 0.000005.
Effigy 4. Quantification of σB-dependent gene expression via RT-qPCR. Fold-change in Log2 calibration of each cistron relative to EDGe is shown for the (A) opuCA; (B) lmo0880; and (C) inlA genes. Each strain was grown at 37 °C with 200 rpm shaking and received either no handling (MQ), osmotic stress (0.5 M NaCl), or sub-MIC (1 µg/mL) norfloxacin. Ct values were normalized to the 16S rRNA factor (ΔCt), and then compared to Border expression (ΔΔCt), and the average was taken from 2 independent experiments. Significance was adamant using a t-exam on the log2 transformed ii − ΔΔCt values. * p < 0.05, ** p < 0.0005, *** p < 0.000005.
Effigy 5. Quantification of σB operon expression via RT-qPCR. Fold-change in Log2 scale of each gene relative to EDGe is shown for the (A) mazF; (B) rsbW; and (C) sigB genes. Each strain was grown at 37 °C with 200 rpm shaking and received either no treatment (MQ), osmotic stress (0.five M NaCl), or sub-MIC (1 µg/mL) norfloxacin. Ct values were normalized to the 16S rRNA gene (ΔCt), and then compared to Edge expression (ΔΔCt), and the average was taken from two independent experiments. Significance was determined using a t-exam on the logtwo transformed 2 − ΔΔCt values. NE: No Expression detected.
Figure 5. Quantification of σB operon expression via RT-qPCR. Fold-change in Log2 scale of each cistron relative to EDGe is shown for the (A) mazF; (B) rsbW; and (C) sigB genes. Each strain was grown at 37 °C with 200 rpm shaking and received either no treatment (MQ), osmotic stress (0.v M NaCl), or sub-MIC (ane µg/mL) norfloxacin. Ct values were normalized to the 16S rRNA gene (ΔCt), and then compared to EDGe expression (ΔΔCt), and the average was taken from two independent experiments. Significance was adamant using a t-test on the logii transformed two − ΔΔCt values. NE: No Expression detected.
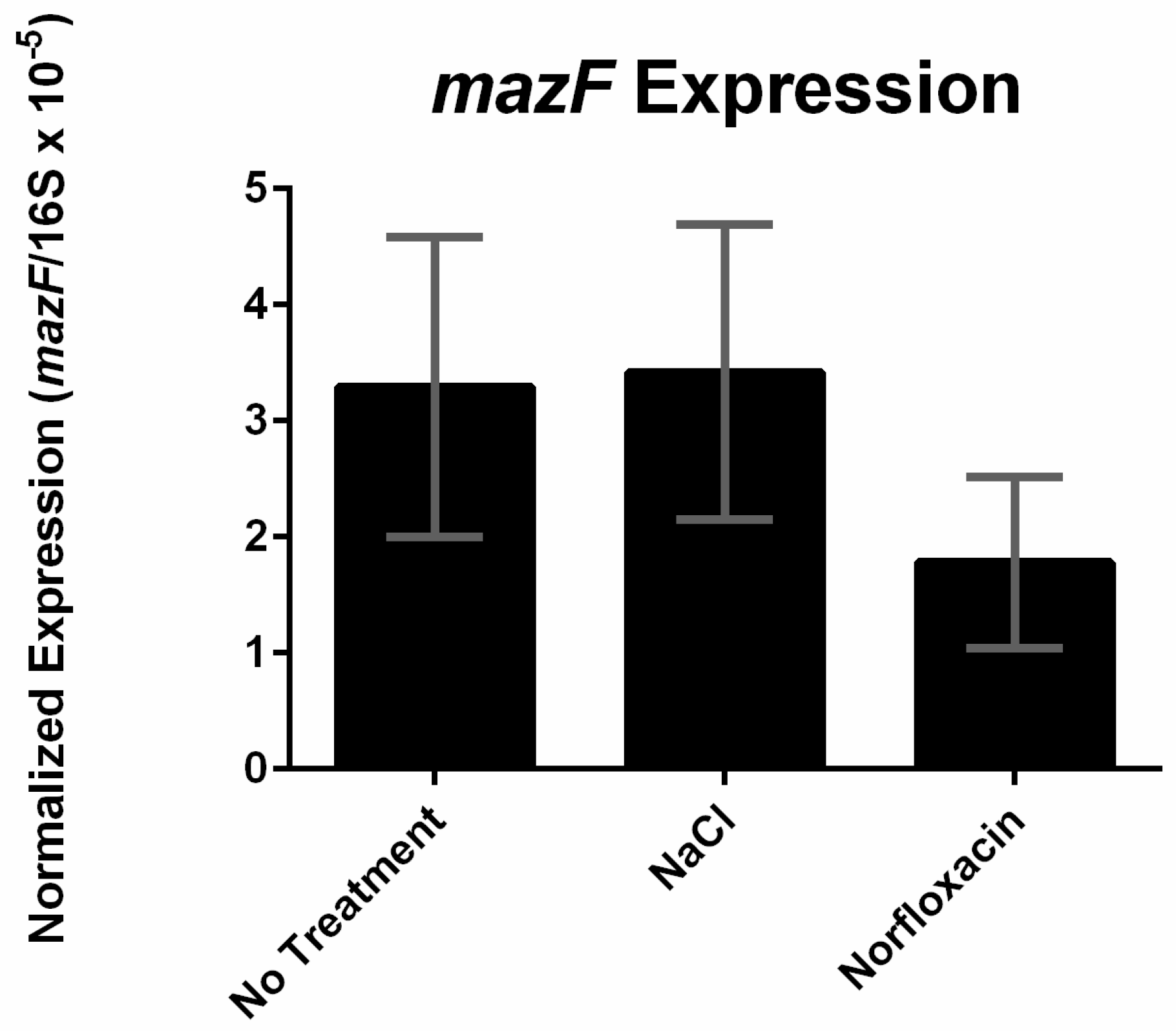
Figure half-dozen. Expression of mazF under different stress conditions (no handling, 0.5 M NaCl, or 1 µg/mL norfloxacin). The re-create number of mazF transcripts were calculated and normalized to the copy number of 16S rRNA transcripts. Error bars represent standard deviation of the mean for 2 biological replicates.
Effigy 6. Expression of mazF nether unlike stress conditions (no treatment, 0.v M NaCl, or 1 µg/mL norfloxacin). The copy number of mazF transcripts were calculated and normalized to the re-create number of 16S rRNA transcripts. Mistake confined represent standard deviation of the mean for two biological replicates.
Tabular array 1. The top x genes with the greatest frequency of MazF cleavage sites (p), every bit determined by comparing the bodily number of MazF cleavage sites to the predicated number based on factor length and nucleotide content.
| Gene Name | Length | UACMU Sites | p | Functional Description |
|---|---|---|---|---|
| lmo2638 | 1887 | 10 | 0.9738 | NADH dehydrogenase |
| secY | 1296 | 8 | 0.9436 | preprotein translocase subunit SecY |
| lmo1310 | 1305 | viii | 0.9413 | hypothetical poly peptide |
| bvrB | 1923 | 8 | 0.9303 | beta-glucoside-specific phosphotransferase enzyme Ii ABC component |
| lmo1911 | 1137 | 7 | 0.8988 | histidine kinase |
| lmo0835 | 1005 | vi | 0.8770 | peptidoglycan binding poly peptide |
| lmo1090 | 984 | 6 | 0.8698 | glycosyltransferase |
| lmo1030 | 1029 | 6 | 0.8600 | LacI family unit transcriptional regulator |
| lmo0880 | 1389 | 6 | 0.8261 | wall associated protein forerunner |
| pgi | 1353 | 6 | 0.8259 | glucose-half-dozen-phosphate isomerase |
Table two. Bacterial strains and plasmids used in this study.
| Strain or Plasmid | Genotype and Relevant Characteristics | Source or Reference |
|---|---|---|
| E. coli DH5α | Plasmid construction and cloning | Lab stock |
| 50. monocytogenes strains | ||
| EGDe | 50. monocytogenes virulent wild-blazon BUG1600, Lineage ΙΙ, serotype 1/2a, MLST ST35 | O. Dussurget |
| ∆clpP | clpP deletion mutant derived from L .monocytogenes EDGe | This report |
| ∆mazEF | lmo0887 and lmo0888 (mazEF antitoxin/toxin) deletion mutant derived from 50. monocytogenes EDGe | This study |
| ∆sigB | sigB deletion mutant derived from L. monocytogenes EDGe | This study |
| Plasmids | ||
| pAUL-A | Temperature sensitive origin of replication, lacZa' multiple cloning site, erythromycin resistance marker | [54] |
Table iii. List of primers used in this study.
| Primer | Purpose | Sequence (5′- > 3′) a,b |
|---|---|---|
| Mutagenesis | ||
| ClpP_UpSt_A | A primer: EcoRΙ | AAAGAATTCCAGTTAATGGGCCAGATT |
| ClpP_UpSt_B | B primer | ACGAATGGTCAAACTAGG |
| ClpP_DnSt_C | C primer | CCTAGTTTGACCATTCGTGCACAAAATGCAAAACCTCT |
| ClpP_DnSt_D | D primer: BamHΙ | AAAGGATCCCGTGACGGATTATTACCA |
| ClpP_IC_Fw | Integration and deletion check | TGGCTCTAACGATGATCTTG |
| ClpP_IC_Rv | TTGATGTTAGTGCACCTGTTG | |
| mazEF_A_Hind | A primer: Hind ΙΙΙ | ATGCAAGCTTTTAGTAGGCGGGGAACTTGCC |
| mazEF_B | B primer | TAACACGTGTCACACCCCCAA |
| mazEF _C | C primer | TTGGGGGTGTGACACGTGTTAGGTTAATGGCTGATGGTGAA |
| mazEF _D_Bam | D primer: BamHΙ | ATGCGGATCCTCAGACCCTTTTGCCCTGC |
| mazEF _IC_Fw | Integration and deletion bank check | CCTTCCACAGAAATCAAAAC |
| mazEF _IC_Rv | CCAACCTTTCTCCACTATT | |
| SigB_A_3 | A primer: EcoRΙ | AAAGAATTCAGCTGTAAGTGAAGCCATCAC |
| SigB_B_3 | B primer | CGCCTCTTTATCAGGTTGAGA |
| SigB_C_3 | C primer | TCTCAACCTGATAAAGAGGCGGTGTCTAGAATCCAACGTCAA |
| SigB_D_3 | D primer: BamHΙ | AAAGGATCCTAATAGCTATCGCAGCACC |
| SigB_IC_Fw | Integration and deletion cheque | CGTCAACGCCAAAGTGAA |
| SigB_IC_Rv | CACCTTTCAAACCATCGCTA | |
| qPCR | ||
| mazF_qPCR_Fw | Quantification of mazF | ACGGCCTGTTCTCATCATTC |
| mazF_qPCR_Rv | expression | CGTTGGCAATTTTGCTTTTT |
| sigB_qPCR_Fw | Quantification of sigB | GAAGCAATGGAAATGGGAAA |
| sigB_qPCR_Rv | expression | CCGTACCACCAACAACATCA |
| rsbW_qPCR_Fw | Quantification of rsbW | ATTACAACTTCCTGCCAAGC |
| rsbW_qPCR_Rv | expression | AATTGCTTCATAAGAAAATCCTG |
| opuCA_qPCR_Fw | Quantification of opuCA | ACATCGATAAAGGAGAATTTC |
| opuCA_qPCR_Rv | expression | GCCGGTTAATCATCTTCATTG |
| 2638_qPCR_Fw | Quantification of lmo2638 | CTGCTGCTACATCTGGTGCT |
| 2638_qPCR_Rv | expression | ACTGGAACCAACCAGGCATA |
| bvrB_qPCR_Fw | Quantification of bvrB | GCAATTGGCGCTAAAACTTC |
| bvrB_qPCR_Rv | expression | ATTGTAACGATGGCGGTTTC |
| 0880_qPCR_Fw | Quantification of lmo0880 | ATTCCGAACAAAATGGCAAA |
| 0880_qPCR_Rv | expression | TTTCTGCAACGGAGACATCA |
© 2017 by the authors; licensee MDPI, Basel, Switzerland. This commodity is an open access article distributed under the terms and conditions of the Creative Eatables Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Source: https://www.mdpi.com/2072-6651/9/1/31/htm
0 Response to "What Sugar Must Be Added to the Growth Media to Ensure Expression of the Pmocab Operon?"
Post a Comment